SEA STAR SKELETAL NETWORKS
2023
The remarkably complex skeletal systems of the sea stars (Echinodermata, Asteroidea) consist of hundreds to
thousands of individual elements (ossicles). This study, for the first time, segmented and analyzed entire skeletals systems of the giant knobby star, Pisaster giganteus. The in-depth
analyses provided a fundamental understanding of the three-dimensional skeletal architecture of the sea star body wall. The high-throughput workflow developed for this purpose, consisted of micro-computed tomography, automated ossicle segmentation, data visualization tools, and additively manufactured models.
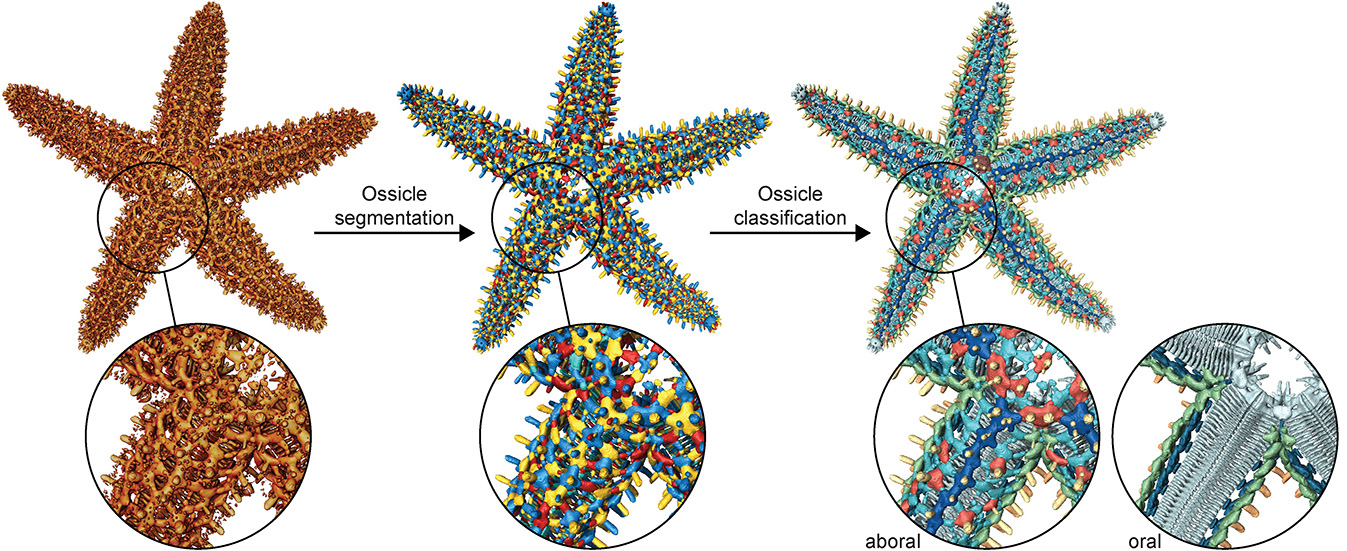
The automated segmentation and ossicle-type classification workflow of a sea star's skeletal system starts with a micro-CT model of the sea star endoskeleton, then individual ossicle segmentation using a random-walk distance-based segmentation method, and lastly color-coded classification based on ossicle type.
With each of the 3,264 ossicles segmented, they were arranged by size and type, allowing us to fully appreciate the sheer complexity of the skeletal system of Pisaster giganteus.
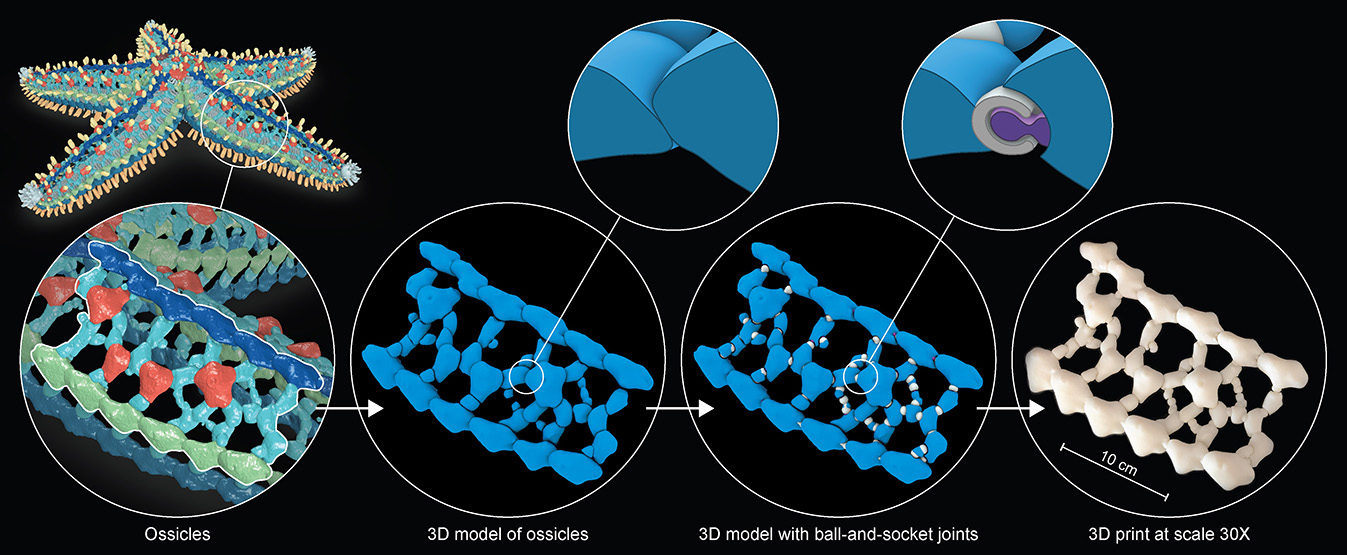
Leveraging the data on ossicle organization obtained through the segmented and classified datasets, fully
articulated 3D-printed models of the skeletal system were generated to explore complex skeletal deformations during arm bending and twisting.
Attribution
Lara Tomholt, Daniel Baum, Rob Wood, James Weaver
Harvard Graduate School of Design
Wyss Institute for Biologically Inspired Engineering at Harvard University
Harvard John A. Paulson School of Engineering and Applied Sciences
Zuse Institute Berlin
Publications
D. Baum, J.C. Weaver, I. Zlotnikov, D. Knötel, L. Tomholt, M.N. Dean. (2019) High-throughput segmentation of tiled biological structures using random-walk distance transforms. Integrative and Comparative Biology, 59 (6), p.1700-1712. https://doi.org/10.1093/icb/icz117
L. Tomholt, D. Baum, R.J. Wood, J.C. Weaver (2023) High-throughput segmentation, data visualization, and analysis of sea star skeletal networks. Journal of Structural Biology, 215 (2), 107955. https://doi.org/10.1016/j.jsb.2023.107955
Media coverage
Press release by Harvard School of Engineering and Applied Sciences - coming soon
Awards
2020Visual Computing for Biology and Medicine (VCBM) Image Contest. Tübingen, Germany. Best Image Award (1st place)
2023Art of Science 2023, Princeton University. Jury Award